Chapter 10: Analysis of Variance: One-way Analysis of Variance
 One-way ANOVA: Using R
One-way ANOVA: Using R
One-Way ANOVA in R
Data and Research Question
The statistical software package #\mathrm{R}# can be used to efficiently conduct a one-way ANOVA. We demonstrate this using the following data, which you can copy and paste into #\mathrm{R}#:
Y <- c(25.6,22.8,27.5,21.3,23.0,33.4,25.2,19.6,29.4,20.8,32.7,23.3,18.2,28.7,24.0)
Group <- c(2,1,3,1,2,3,2,1,3,1,3,2,1,3,2)
Data <- data.frame(Y, Group)
Here #\mathtt{Y}# is the outcome variable, and #\mathtt{Group}# is the categorical factor, which has 3 levels. There are #15# observations, and the #k#th position in the two vectors gives the level of the factor and the value of the outcome variable for the #k#th subject, for #k = 1,...,15#.
We merge these together into a data frame that we conveniently name #\mathtt{Data}#. (If you read data into #\mathrm{R}# from a file using the #\mathtt{read.table()}# function, it will be loaded as a data frame. You can give it any name you like, though Data usually makes the most sense.)
The research question we would like to answer is: Is the population mean of the outcome variable #Y# the same across all three groups?
One-way Analysis of Variance
To test the null hypothesis of equal population means, we first create the ANOVA model using the #\mathtt{aov()}# function.
> Model <- aov(Y ~ factor(Group), data = Data)
We store the output from the #\mathtt{aov()}# function in an object that we name #\mathtt{Model}#. You can choose any name you like.
Note also that we have to convert the numeric vector #\mathtt{Group}# into the factor class using the #\mathtt{factor()}# function if we want the #\mathtt{aov() }# function to work properly. Even if #\mathtt{Group}# was already in the factor class, using the #\mathtt{factor()}# function would do no harm, so to be on the safe side it is a good idea to always do this.
Note also that the #\mathtt{aov()}# function allows us to specify the data frame in which to find the variables, so that we don't have to type #\mathtt{Data$Y}# and #\mathtt{Data$Group}#. If the variables are not nested inside a data frame, but are in the #\mathrm{R}# workspace, then we would not use the "#\mathtt{data=}#" setting.
To inspect the sample means along with the grand sample mean, we use the #\mathtt{model.tables()}# function on the #\mathtt{Model}# object, with the argument #\mathtt{type=}# "#\mathtt{means}#":
> model.tables(Model, type = "means")
Tables of means
Grand mean
25.03333
factor(Group)
factor(Group)
1 2 3
20.54 24.22 30.34
Now that the ANOVA model has been constructed, we next use the #\mathtt{summary()}# function on the #\mathtt{Model}# object to inspect the results of the analysis:
> summary(Model)
Df Sum Sq Mean Sq F value Pr(>F)
factor(Group) 2 245.06 122.53 33.47 1.23e-05 ***
Residuals 12 43.93 3.66
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
The output of the #\mathtt{summary()}# function is an ANOVA table. Here the given #p#-value is #p=0.0000123#, which is between #0# and #0.001#, as signified by the #3# stars next to the value.
As such, we would reject the null hypothesis and conclude that there are at least two population means which differ out of the three groups. However, this conclusion is only valid if the assumptions are satisfied.
Checking the Assumptions
If the conclusion of the one-way ANOVA is to reject the null hypothesis of equal means, then there are two assumptions that we need to check before moving on to the post hoc analysis:
- The assumption of normality is checked by determining whether or not the residuals of the model are normally distributed.
- The homogeneity of variance assumption is checked by determining whether or not the variance of the residuals is the same across all groups.
In both cases, we first plot the relevant data to get a first impression of the situation, before proceeding with a formal test of the assumption.
In order to visually inspect the normality of the outcome variable #\mathtt{Y}# in each group, we create a Q-Q-plot of the model residuals:
> qqnorm(Model$residuals, pch=20)
> qqline(Model$residuals)
The first function #\mathtt{qqnorm()}# draws the scatterplot and the second function #\mathtt{qqline()}# creates the reference line. This creates the following Q-Q-plot: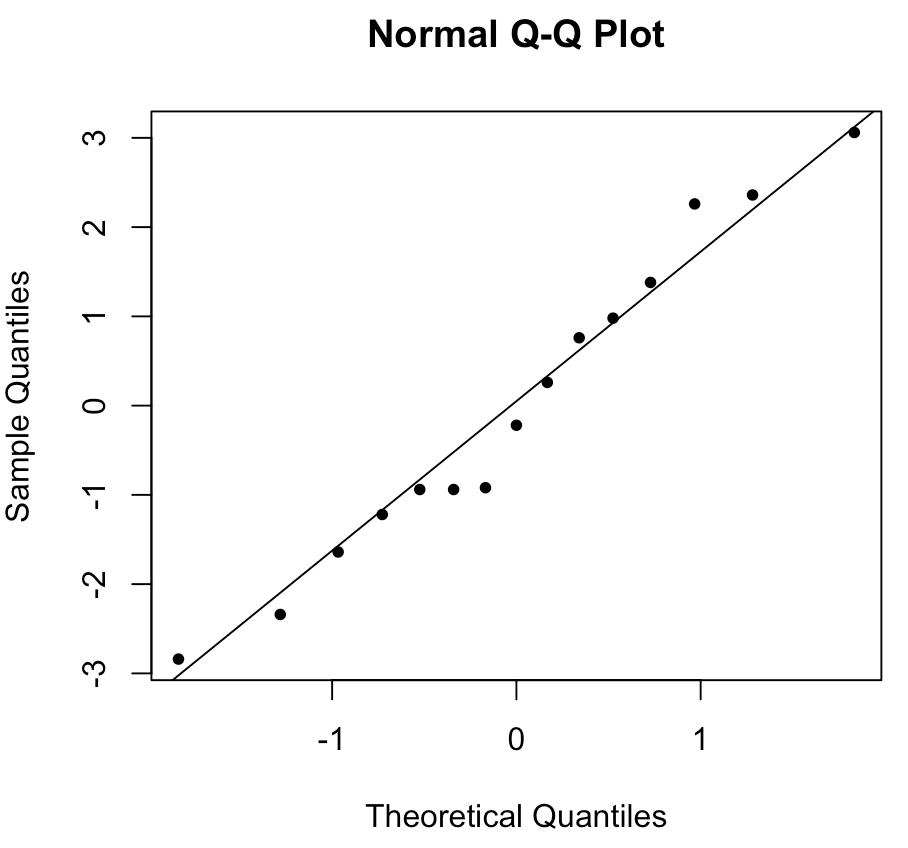
In this case, the points are close enough to the line that we can be fairly confident that the assumption of normality is satisfied.
To make sure the assumption of normality is not violated, we next run the Shapiro-Wilk test to formally test the following hypotheses:
\[\begin{array}{rcl}
H_0&:&\text{The residuals are normally distributed.}\\\\
H_a&:&\text{The residuals are not normally distributed.}
\end{array}\] To run the test, we use the #\mathtt{shapiro.test()}# function on the residuals of the #\mathtt{Model}# object:
> shapiro.test(Model$residuals)
Shapiro-Wilk normality test
data: Model$residuals
W = 0.96597, p-value = 0.7945
Here, the #p#-value of #p=0.7945# is so large that the null hypothesis of the test would not be rejected at any reasonable significance level #\alpha#. We can thus conclude that the normality assumption is satisfied.
In order to visually inspect the variance of the residuals, we make a scatterplot of the model residuals against the fitted values (i.e., the group sample means):
> plot(Model$fitted.values, Model$residuals, pch=20, xlab="Fitted values", ylab="Residuals")
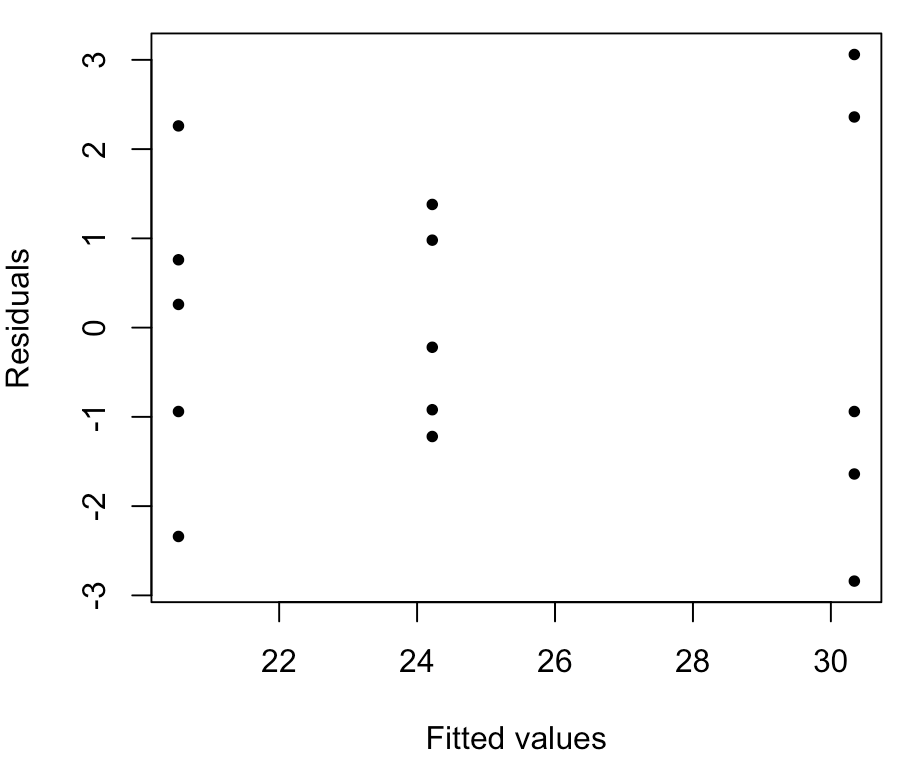
Here, the spread of the points in the middle group (group 2) is noticeably smaller than the spread of the points in the other two groups, which indicates that we may have a problem with the equal variance assumption. But this is hard to address visually.
To formally assess the homogeneity assumption we use Bartlett's test to test the following hypotheses:
\[\begin{array}{rcl}
H_0&:&\text{The population variances are all equal.}\\\\
H_a&:&\text{The population variances are not all equal.}
\end{array}\] To run the test, we use the #\mathtt{bartlett.test()}# function, which takes the same input as the #\mathtt{aov()}# function we used earlier:
> bartlett.test(Y ~ factor(Group), data = Data)
Bartlett test of homogeneity of variances
data: Y by factor(Group)
Bartlett's K-squared = 2.2327, df = 2, p-value = 0.3275
Here, the #p#-value of #p=0.3275# is so large that the null hypothesis of the test would not be rejected at any reasonable significance level #\alpha#. We can thus conclude that the homogeneity of variance assumption is satisfied.
Post Hoc Analysis
The final step of the ANOVA procedure is to run a post hoc analysis to determine which of the population means differ from each other.
Since the assumption of equal population variances is satisfied, the Tukey-Kramer HSD test is recommended. To run this test, we use the #\mathtt{TukeyHSD()}# function, which takes the #\mathtt{Model}# object as its input:
> TukeyHSD(Model)
Tukey multiple comparisons of means
95% family-wise confidence level
Fit: aov(formula = Y ~ factor(Group), data = Data)
$`factor(Group)`
diff lwr upr p adj
2-1 3.68 0.4515543 6.908446 0.0257595
3-1 9.80 6.5715543 13.028446 0.0000092
3-2 6.12 2.8915543 9.348446 0.0007588
Using a significance level of #\alpha = 0.01#, we would conclude that there exists a significant difference between the population means of groups #1# and #3# and between the means of groups #2# and #3#, but not between the means of groups #1# and #2#.

Or visit omptest.org if jou are taking an OMPT exam.



